|
1/7/2024 0 Comments Box plots in igor pro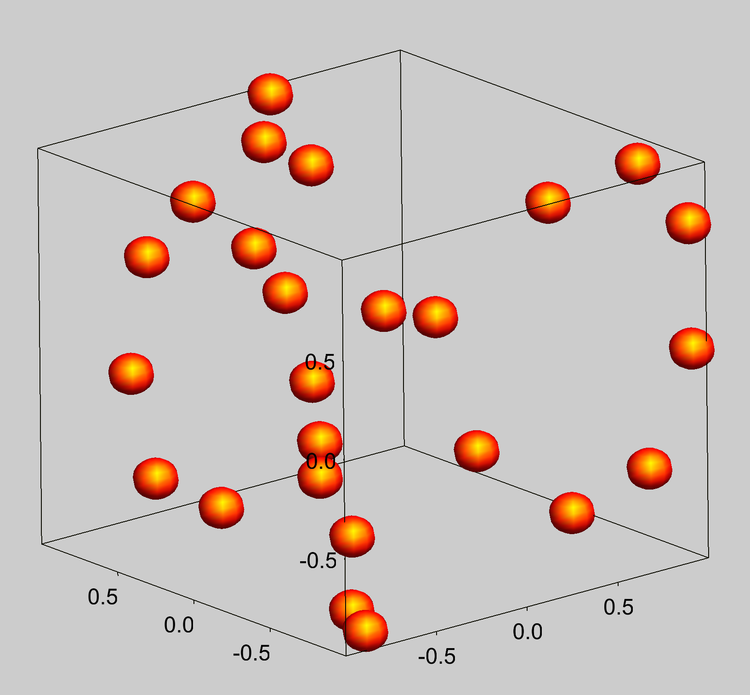
Our work indicates that while both early and modern amino acids are predisposed to supporting protein structure, they do so with different biophysical properties and via different mechanisms. By contrast, chaperones significantly increase the otherwise poor solubility of the modern alphabet proteins suggesting their coevolution with the amino acid repertoire. In addition, the early alphabet proteins are inherently more soluble and refoldable, independent of the general Hsp70 chaperone activity. We show that compact conformations resistant to proteolysis are surprisingly similarly abundant in both libraries. The first is composed of the entire alphabet of 20 amino acids while the second one consists of only 10 residues (ASDGLIPTEV) representing a consensus view of plausibly available amino acids through prebiotic chemistry. 
Here, we analysed two combinatorial protein libraries representing proxies of the available sequence space at two different evolutionary stages. How (or if) early proteins could sustain an early biosphere has been a major puzzle. In extant proteins, a significant fraction of the ‘late’ amino acids (such as Arg, Lys, His, Cys, Trp and Tyr) belong to essential catalytic and structure-stabilizing residues. The earliest proteins had to rely on amino acids available on early Earth before the biosynthetic pathways for more complex amino acids evolved. 
We discuss the case of sulfur-containing amino acids as a specific and clear example and conclude by reviewing the potential impact of better understanding the chemical “logic” of a smaller forerunner to the standard amino acid alphabet. Findings support the idea that a small sample size of amino acids caused previous ambiguous results, and further improvements in meteorite analysis, and/or prebiotic simulations will further clarify the nature and extent of unusual properties. However, the strength of this pattern varies depending on both the size and composition the library used to create a background (null model) for a random alphabet, and the precise definition of exactly which amino acids were present in a simpler, earlier code. 
In addition to confirming the well-established idea that “late” arriving amino acids expanded the chemistry space encoded by genetic material, we find that a prebiotically plausible subset of coded amino acids generally emulates the patterns found in the full set of 20, namely an exceptionally broad and even distribution of volumes and an exceptionally even distribution of hydrophobicities (quantified as logP) over a narrow range. Here, we test this hypothesis using significantly updated data for organic material detected within meteorites, which contain several coded and non-coded amino acids absent from prior studies. We have suggested previously that this ambiguity may result from the low number of amino acids recognized by the definition of prebiotic plausibility used for the analysis. These findings motivate a re-examination of prior work which has identified unusual properties of the set of twenty amino acids found within the full genetic code, while leaving it unclear whether similar patterns also characterize the subset of prebiotically plausible amino acids. Recent findings, in vitro and in silico, are strengthening the idea of a simpler, earlier stage of genetically encoded proteins which used amino acids produced by prebiotic chemistry.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
